A research team led by Professor Yongjin Zhou from the Dalian Institute of Chemical Physics, Chinese Academy of Sciences, together with Associate Professor Hongzhong Lu from Shanghai Jiao Tong University, published their findings in Proceedings of the National Academy of Sciences (PNAS). They developed a novel systems biology analysis method specifically for Saccharomyces cerevisiae (brewing yeast) strains. This method reveals the adaptive mechanisms of yeast in different environments by constructing strain-specific metabolic models.
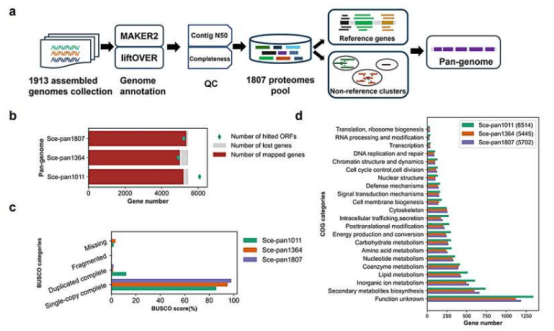
The research team established a high-quality digital resource of the yeast pangenome and constructed metabolic models tailored to individual strains. They developed an integrated analysis pipeline that combines genomic data, metabolic models, and multi-omics information, enabling systematic evaluation of the characteristics of different yeast strains. Currently, research on Saccharomyces cerevisiae is mostly limited to a few laboratory strains such as S288c and CEN.PK, which restricts the discovery of optimal strains for developing high-performance cell factories.
To validate the method, the researchers applied it to the analysis of industrial ethanol-producing strains. The study found that enhancing the downstream pathways of glycolysis is key to achieving efficient ethanol synthesis, a finding validated at the genetic, transcriptional, and metabolic levels. This research provides new ideas for the rational design of yeast cell factories used in the production of ethanol and its derivatives.
Professor Yongjin Zhou stated: "Our work not only provides the academic and industrial communities with a comprehensive digital resource of yeast strains, but also offers a new method for evaluating and selecting the best chassis strains for biomanufacturing applications." This method is expected to promote broader applications of yeast strains in the biotechnology field.
