Cucumbers are not only a common ingredient in summer salads and sandwiches, and a valuable economic crop, but also a model plant that helps researchers push the boundaries of genomic discovery. Recently, a collaborative study between the UK’s John Innes Centre and the Chinese Academy of Agricultural Sciences (CAAS) has achieved a major breakthrough in cucumber gene research. The results were published in the journal Cell.
This collaborative study used a series of experiments and genomic analyses to deeply investigate the differences between wild cucumbers and their domesticated relatives at the molecular level, focusing on the genes responsible for the elongation of domesticated cucumber fruit. Compared to their short, bitter wild relatives, domesticated cucumbers have longer fruit—a trait that has long been a mystery the research team sought to unravel.
Modern plant breeding has largely focused on mutations in DNA sequences that code for proteins, as proteins act as cellular machinery that transmits traits such as fruit length, taste (bitter or sweet), and seed shape (round or wrinkled). However, protein-coding genes account for only a small portion of the genome. Increasingly, researchers are exploring non-coding DNA sequences, where synonymous mutations (previously called silent mutations) in these regions are gradually gaining attention from biologists. Although previous studies have suggested that synonymous mutations play roles in cellular function, there has been little evidence that they can shape biological traits in multicellular organisms.
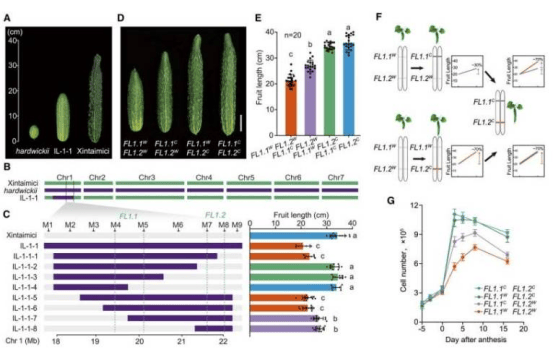
In this study, leveraging a population-based genomic variation map of cucumbers, the research team identified fruit length as a key domestication trait. Through molecular and genetic analyses, they revealed the precise mechanism behind cucumber elongation: a synonymous mutation (1287C>T) in the ACS2 gene was the key driver of fruit elongation during cucumber domestication. This mutation eliminates an m⁶A RNA modification site, reshapes the RNA structure to suppress translation, reduces ACS2 translation and ethylene levels, and ultimately increases fruit length by up to 70%.
Notably, this mutation does not lead to changes in protein production like most agriculturally and biologically important traits; instead, it acts on another molecule—RNA—reshaping it to suppress the production of the protein responsible for the “short” trait in wild cucumbers. Dr. Zhang Yueying, a postdoctoral researcher at the John Innes Centre and first author of the study, said: “A tiny ‘silent’ change in the cucumber gene, long considered harmless, turned out to be the key factor that made modern cucumbers grow longer. This long-assumed biologically neutral silent mutation reconnected RNA regulation and directly promoted the development of domestication traits.”
These findings provide valuable insights for crop breeding programs and offer potential pathways for future trait gene engineering, particularly for key traits such as fruit size that can increase crop yield and bring commercial benefits to growers. At the same time, the study paves the way for further research targeting synonymous sites and holds promise for using precise crop improvement technologies such as gene editing to enhance field traits in various crops.
The study was completed through collaboration between the team led by Professor Ding Yiliang at the John Innes Centre (UK), the team led by Professor Yang Xueyong at the Institute of Vegetables and Flowers of the Chinese Academy of Agricultural Sciences, and Academician Huang Sanwen, President of the Chinese Academy of Agricultural Sciences.
