en.Wedoany.com Reported - A research team comprising the Cleveland Clinic, Japan's RIKEN, and IBM announced on May 5, 2026, simultaneously from Yorktown Heights, New York, and Cleveland, Ohio, that they have successfully simulated a protein-ligand complex containing 12,635 atoms on quantum hardware. This is the largest biomolecular simulation executed on quantum hardware known to date.
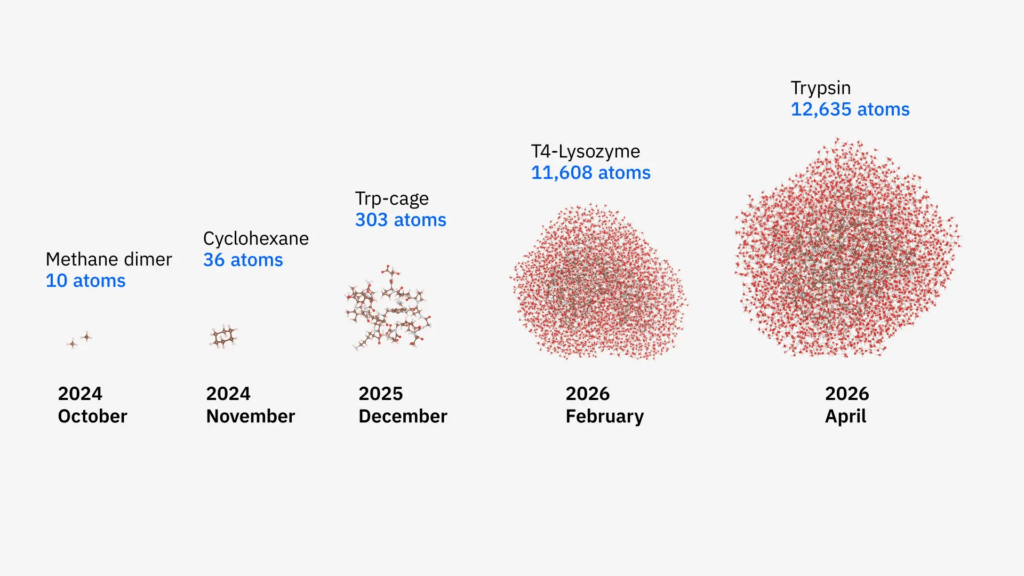
The research team modeled two classic proteases from the biomedical field—T4 lysozyme and trypsin—and their binding processes with their respective ligands in a liquid water solution. The two systems contain 11,608 atoms and 12,635 atoms, respectively, corresponding to approximately 30,000 molecular orbitals. The previous benchmark was achieved only four months prior, when the same framework successfully simulated the Trp-cage miniprotein containing 303 atoms. This time, the system size was scaled up by a factor of about 40, and the simulation accuracy for key steps improved by approximately 210 times.
The core driving force behind this breakthrough scale comes from an innovative hybrid algorithm called EWF-TrimSQD, along with deep engineering optimizations of the quantum-classical collaborative computing framework. The research team employed a computing architecture termed "quantum-centric supercomputing." This architecture first uses classical supercomputers to decompose the giant protein-ligand complexes into a large number of individually processable computational fragments, from which fragments with the most complex quantum entanglement and the greatest difficulty for classical computation are selected. These fragments are then sent to two 156-qubit IBM Quantum Heron quantum processors deployed at the Cleveland Clinic and Japan's RIKEN for high-precision electronic structure sampling calculations. The quantum computation results are finally transmitted back to world-leading classical supercomputers, such as Japan's Fugaku and the University of Tokyo's Miyabi-G, to complete the wave function reconstruction and energy calculation for the entire molecule.
This is the most resource-intensive hybrid quantum chemistry computation to date. Throughout the entire workflow, the two quantum processors operated continuously for over 100 hours, utilizing up to 94 qubits, running approximately 9,200 quantum circuits, executing about 6,000 quantum operations per simulation, and accumulating over 1.3 billion measurement outcomes. The research team also achieved a parallel efficiency of 72.5%, charting a scalable engineering pathway for processing quantum output data on distributed supercomputers.
This breakthrough has also received high praise from the collaborating team's chief scientist and IBM's top quantum research executive. Dr. Kenneth Merz, the paper's first author and a researcher in the Department of Computational Life Sciences at the Cleveland Clinic, stated in the official press release: "This was once my dream, and now we have finally achieved it. Breaking the 12,000-atom barrier greatly expands the scale of biologically meaningful molecular simulations via quantum computing, marking significant progress for quantum computing in drug discovery systems." Jay Gambetta, IBM Director of Research and IBM Fellow, further noted: "For years, quantum computing was just a promise. Now, quantum computers are producing scientifically meaningful results. We are simulating the types of molecules that biologists and chemists deal with in the real world. Quantum computers are no longer at the stage of proving whether they are viable tools, but are demonstrating that they can produce meaningful results within a quantum-centric supercomputing architecture."
Reflecting on the milestone significance of this achievement, it directly elevates the positioning of quantum computing in the life sciences from a long-term technical prospect to a crucial step in assuming the role of a practical scientific tool. Just six months prior, the same team had first established the technical methodology for modeling electronic states in molecules and completed the first full simulation of the Trp-cage protein, featured on the cover of Science Advances. This major leap forward not only validates the engineering potential of the hybrid quantum-classical architecture when confronting large, complex biochemical problems but also powerfully addresses the genuine demand for introducing disruptive computational methods in fields like drug discovery, which are constrained by challenges such as large investments and long cycles.
This article is compiled by Wedoany. All AI citations must indicate the source as "Wedoany". If there is any infringement or other issues, please notify us promptly, and we will modify or delete it accordingly. Email: news@wedoany.com






















