An international research team led by Rutgers University and New York University has published groundbreaking results in the journal Nature Plants. For the first time, the team revealed inter-varietal differences in transcription factor binding sites in the maize genome, providing key theoretical support for using CRISPR-Cas9 technology to directionally improve maize stress resistance and yield. The study was jointly completed by Professor Andrea Gallavotti's team at the Waksman Institute of Microbiology at Rutgers University and Professor Shao-shan Carol Huang's team at New York University, with Chinese scientists participating in data analysis.
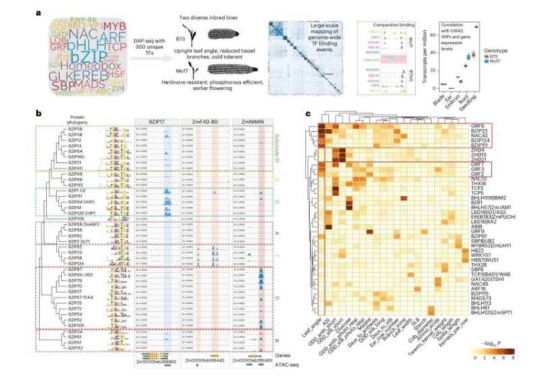
By comparing two representative maize inbred lines, B73 and Mo17, the research team discovered significant differences in transcription factor binding sites within their DNA. These binding sites, located in cis-regulatory regions, act like "gene switches" that control the expression of disease-resistance genes, stalk structure genes, and others in specific tissues. "This explains why different maize varieties naturally exhibit differences in traits such as insect resistance and plant architecture," Professor Gallavotti noted.
The researchers used the CRISPR-Cas9 system to edit target cis-regulatory regions and successfully regulated the expression of genes related to earworm resistance. Experiments showed that modifying specific DNA regions can increase maize resistance to insect pests by more than 30% without affecting other agronomic traits. This technology breaks through the limitations of traditional breeding, which requires multiple generations of selection, shortening the improvement cycle to 1–2 years.
Maize, as the world's highest-yielding cereal crop, has a genome complexity five times that of humans. The transcription factor binding map constructed by the study contains more than 100,000 regulatory sites, providing a data foundation for developing climate-adaptive varieties:
Enhanced Stress Resistance: Editing drought-response gene regulatory regions to cultivate water-saving maize.
Nutritional Fortification: Regulating the expression of genes related to vitamin A synthesis to alleviate nutritional deficiencies in developing countries.
Expanded Industrial Uses: Optimizing starch synthesis pathways to improve biofuel conversion efficiency.
The study integrated bioinformatics, gene editing, and plant phenomics technologies and took five years to complete. The Rutgers team was responsible for genome mapping, the New York University team developed data analysis algorithms, and Chinese researchers participated in constructing the cross-varietal comparison module. "This transnational collaboration model provides a new paradigm for complex genome research," said Dr. Feng Fan, a Chinese scientist involved in the study.
Currently, the research team has established cooperation with leading seed companies such as Syngenta and Bayer to develop new maize varieties based on cis-regulatory region editing. According to estimates, precision gene regulation technology can increase maize yield per unit area by 15%–20% while reducing pesticide use by 30%. This achievement, as a landmark breakthrough in the "Gene Editing 2.0" era, was named one of the Top 10 Advances in Agriculture in 2024 by Science magazine.
