A team of scientists from the University of New South Wales (UNSW) has developed a new artificial intelligence tool called ECSFinder, designed to decode the 98% of the human genome that consists of non-coding regions. The study, conducted in collaboration with the University of Montreal and McGill University in Canada, was published in the journal Nucleic Acids Research.
Only about 2% of the human genome is responsible for encoding proteins, while the rest has long been regarded as "junk DNA." Associate Professor Martin Smith from the School of Biotechnology and Biomolecular Sciences at UNSW said: "We are trying to decode the logic circuits of the human genome — the hidden rules that tell our DNA how to build and run a human." Research indicates that the roots of many diseases, including heart disease, cancer, and certain mental disorders, may lie in these non-coding regions.
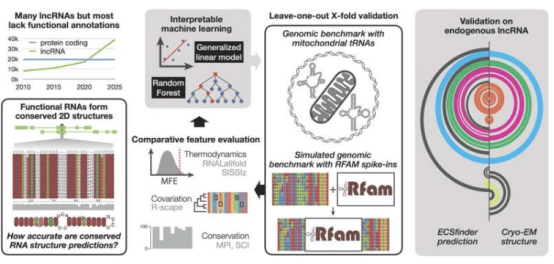
The ECSFinder tool explores genome function by identifying RNA structures. The research team trained an artificial intelligence system using known RNA structures, enabling it to detect hidden regulatory elements in the genome. "We expect to discover hundreds of thousands of new RNA structures, adding a new dimension to our understanding of the genome," Associate Professor Smith added. The tool demonstrated superior performance compared to existing methods in testing.
The non-coding regions of the genome contain long non-coding RNAs, which have critical regulatory functions and can activate or inhibit the expression of specific genes. Associate Professor Smith explained: "We believe these RNAs act like software, coordinating the protein 'hardware' into a normally functioning symphony." Understanding this regulatory mechanism will provide new insights for disease treatment.
The researchers noted that the conserved nature of non-coding regions in the genome indicates their important biological functions. Although humans have diverged evolutionarily from organisms such as bananas, certain genome sequences remain highly conserved. The application of the ECSFinder tool will help discover regulatory elements that are difficult to detect with traditional methods.
The research team believes that a deeper understanding of the non-coding regions of the genome will drive the development of personalized medicine. Associate Professor Smith stated: "Since RNA structures can serve as targets for drugs, they offer an exciting new frontier for treatment." By decoding the non-coding regions of an individual's genome, more precise treatment strategies may be designed for specific diseases in the future.
